TLDR: The number of subsampled features is a main source of randomness and an important parameter in random forests. Mind the different default values across implementations.
Randomness in Random Forests
Random forests are very popular machine learning models. They are build from easily understandable and well visualizable decision trees and give usually good predictive performance without the need for excessive hyperparameter tuning. Some drawbacks are that they do not scale well to very large datasets and that their predictions are discontinuous on continuous features.
A key ingredient for random forests is—no surprise here—randomness. The two main sources for randomness are:
- Feature subsampling in every node split when fitting decision trees.
- Row subsampling (bagging) of the training dataset for each decision tree.
In this post, we want to investigate the first source, feature subsampling, with a special focus on regression problems on continuous targets (as opposed to classification).
Feature Subsampling
In his seminal paper, Leo Breiman introduced random forests and pointed out several advantages of feature subsamling per node split. We cite from his paper:
The forests studied here consist of using randomly selected inputs or combinations of inputs at each node to grow each tree. The resulting forests give accuracy that compare favorably with Adaboost. This class of procedures has desirable characteristics:
i Its accuracy is as good as Adaboost and sometimes better.
ii It’s relatively robust to outliers and noise.
iii It’s faster than bagging or boosting.
iv It gives useful internal estimates of error, strength, correlation and variable importance.
v It’s simple and easily parallelized.
Breiman, L. Random Forests. Machine Learning 45, 5–32 (2001).
Note the focus on comparing with Adaboost at that time and the, in today’s view, relatively small datasets used for empirical studies in this paper.
If the input data as p number of features (columns), implementations of random forests usually allow to specify how many features to consider at each split:
| Implementation | Language | Parameter | Default |
| scikit-learn RandomForestRegressor | Python | max_features | p |
| scikit-learn RandomForestClassifier | Python | max_features | sqrt(p) |
| ranger | R | mtry | sqrt(p) |
| randomForest regression | R | mtry | p/3 |
| randomForest classification | R | mtry | sqrt(p) |
| H2O regression | Python & R | mtries | p/3 |
| H2O classification | Python & R | mtries | sqrt(p) |
Note that the default of scikit-learn for regression is surprising because it switches of the randomness from feature subsampling rendering it equal to bagged trees!
While empirical studies on the impact of feature for good default choices focus on classification problems, see the literature review in Probst et al 2019, we consider a set of regression problems with continuous targets. Note that different results might be more related to different feature spaces than to the difference between classification and regression.
The hyperparameters mtry, sample size and node size are the parameters that control the randomness of the RF. […]. Out of these parameters, mtry is most influential both according to the literature and in our own experiments. The best value of mtry depends on the number of variables that are related to the outcome.
Probst, P. et al. “Hyperparameters and tuning strategies for random forest.” Wiley Interdisciplinary Reviews: Data Mining and Knowledge Discovery 9 (2019): n. pag.
Benchmarks
We selected the following 13 datasets with regression problems:
| Dataset | number of samples | number of used features p |
| Allstate | 188,318 | 130 |
| Bike_Sharing_Demand | 17,379 | 12 |
| Brazilian_houses | 10’692 | 12 |
| ames | 1’460 | 79 |
| black_friday | 166’821 | 9 |
| colleges | 7’063 | 49 |
| delays_zurich_transport | 27’327 | 17 |
| diamonds | 53’940 | 6 |
| la_crimes | 1’468’825 | 25 |
| medical_charges_nominal | 163’065 | 11 |
| nyc-taxi-green-dec-2016 | 581’835 | 14 |
| particulate-matter-ukair-2017 | 394’299 | 9 |
| taxi | 581’835 | 18 |
Note that among those, there is no high dimensional dataset in the sense that p>number of samples.
On these, we fitted the scikit-learn (version 0.24) RandomForestRegressor (within a short pipeline handling missing values) with default parameters. We used 5-fold cross validation with 4 different values max_feature=p/3 (blue), sqrt(p) (orange), 0.9 p (green), and p (red). Now, we show the mean squared error with uncertainty bars (± one standard deviation of cross validation splits), the lower the better.
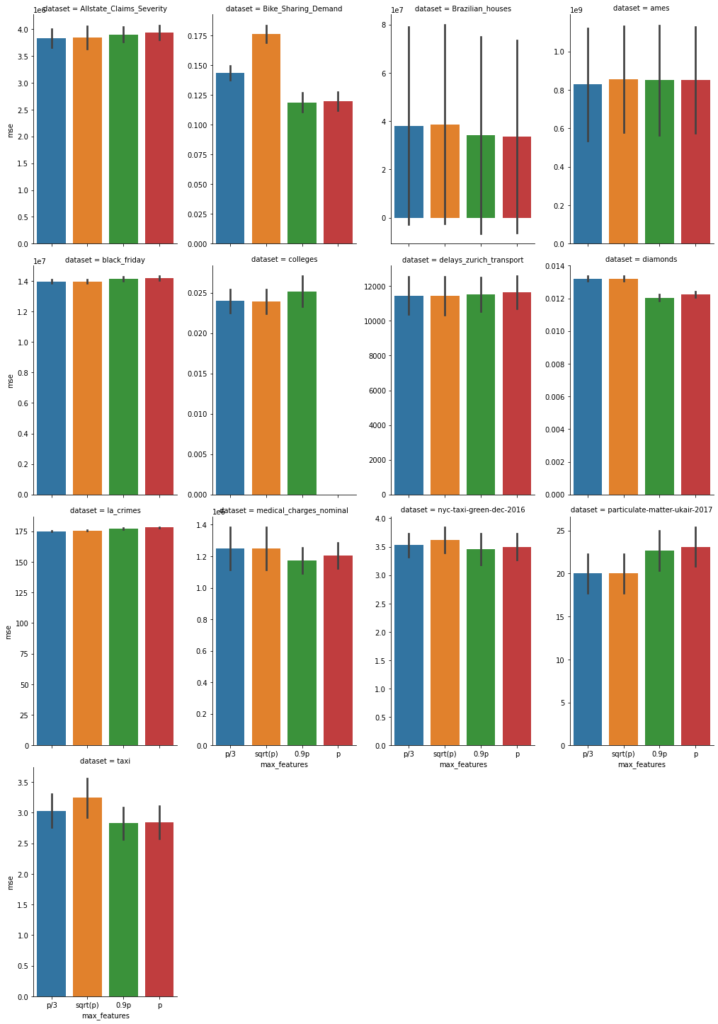
In addition, we also report the fit time of each (5-fold) fit in seconds, again the lower the better.
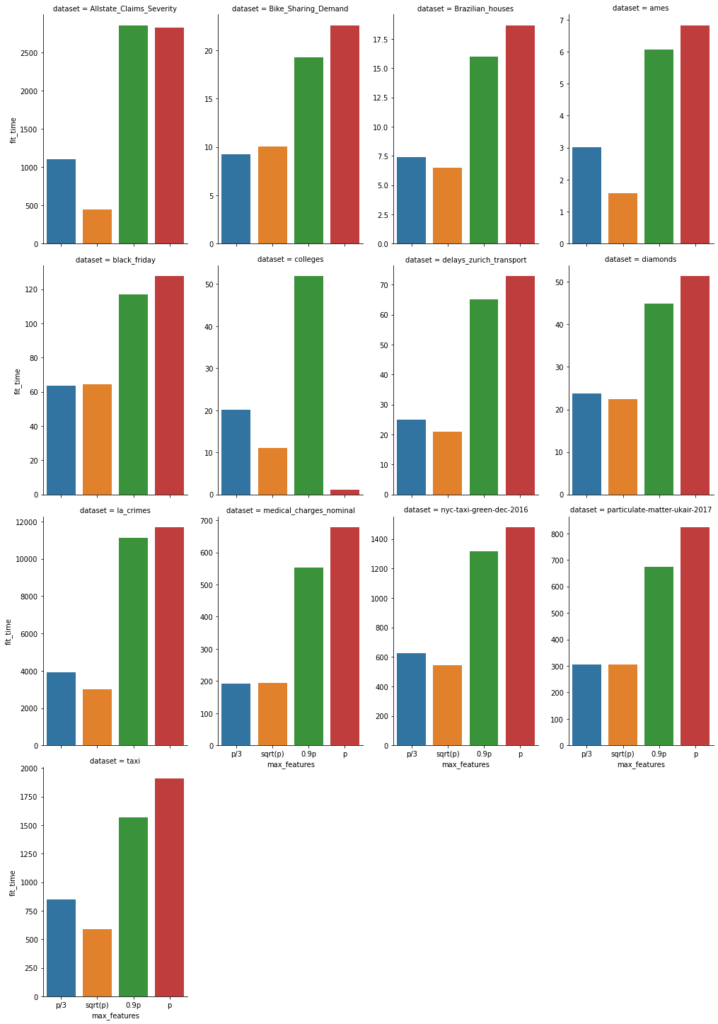
Note that sqrt(p) is often smaller than p/3. With this in mind, this graphs show that fit time is about proportional to the number of features subsampled.
Conclusion
- The main tuning parameter of random forests is the number of features used for feature subsampling (max_features, mtry). Depending on the dataset, it has a relevant impact on the predictive performance.
- The default of scikit-learn’s RandomForestRegressor seems odd. It produces bagged trees. This is a bit like using ridge regression with a zero penalty😉. However, it can be justified by our benchmarks above.
The full code can be found here: